Pores and Selectivity Filters
In this post I will be briefly introducing the classical ideas of ion conduction through membrane proteins. The idea of ion channels as selective pores in the cell membrane is very old but is now understood at atomic detail. So what is a channel? What does an ion conductive pore look like? Here, I will use the structures of potassium selective ion channels to briefly discuss these questions.
All cells are surrounded by a membrane. These membranes separate the contents inside the cell (in the cytoplasm) from the outside. The membrane is made up of amphipathic lipid molecules that form a hydrophobic barrier which prevents the diffusion of charged and polar molecules into and out of the cell. However, in order to get nutrients from the environment and to dispose of metabolic waste products, it is necessary for the cell to allow certain molecules and ions in and out. A class of membrane spanning proteins have evolved to catalyze the transport or conduction of these small molecules and ions. Ion channels are a specific group of these membrane spanning proteins characterized by two major biophysical properties: gating and selectivity.
Gating is the process by which the channels open and close. Mechanisms of gating vary, for example, some channels are gated by changes in the electrical potential across the membrane (voltage-gated channels) while others are gated by the binding of a small molecule or ion to a regulatory domain (ligand-gated channels).
Selectivity is a preference of the channel to conduct a specific ion. There are five major classes of selective ion channels: proton (H+), sodium (Na+), potassium (K+), calcium (Ca+) and chloride (Cl–). The degree of selectivity can vary between different members of the same class and there are also many types of non-selective channels. Furthermore, there are many channels that conduct other less abundant ions. The properties of ion channels that make them selective are still not totally understood.
The most well understood example of specific conduction of ions across the membrane was revealed to us by the structures of the K+ selective channels. In this post I will briefly explore two basic features of these structures that help us understand how ions traverse the membrane and how channels can select between different ionic species (i.e. K+ vs Na+).
—
Pores
Charged atoms and molecules cannot easily traverse the cell membrane. The hydrophobic interior of the membrane is unable to stabilize the charge of the ion whereas polar water molecules can orient around the ion stabilizing its charge. This basically means that ions stay in water and are unable to penetrate the cell membrane. So how do ion channels make it easier for the ions to cross the membrane? These proteins span the membrane and generate water filled pores that allow water molecules to stabilize the ions as they pass through the membrane. In addition, the polar properties of the amino acid residues making up the protein main chain, side chains and secondary structure (helix dipoles) contribute to stabilizing the ions (Gouaux & MacKinnon, 2005). Fig.1 shows the open pore of the voltage-gated K+ channel (Long et al. 2007).
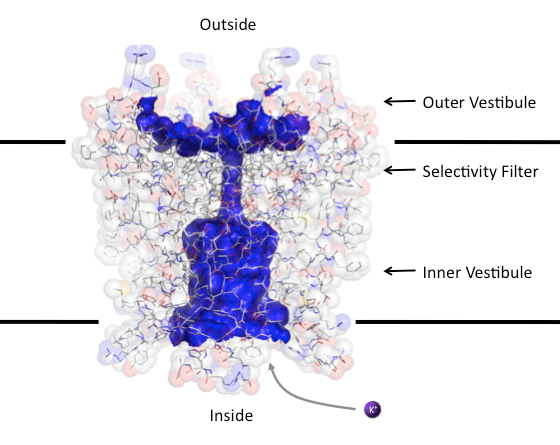
Figure 1. Open pore of the voltage-gated potassium channel. Blue surface coats the water accessible channel pore. The amino acids of the channel are shown as sticks surrounded by transparent space-filling spheres. The residues are colored by atom with carbon grey, oxygen red, nitrogen blue and sulfur yellow. The black lines indicate the approximate boundaries of the membrane. The surface of the pore was calculated using the program Hollow and the figure was made using PyMol.
Hydrated ions can simply diffuse into the pore and cross the membrane. In general ion channels allow for the passive diffusion of ions down their concentration gradient with no strong preference whether the ion is going into or out of the cell. Therefore, even though in Fig.1 I show a K+ ion entering the channel from the inside of the cell (leaving the cell) it could just as easily be diffusing into the cell from the outside. When the channel is open the rate limiting step for the conduction of the ion is diffusion into the pore (Morais-Cabral et al., 2001).
The narrowest part of the conduction pathway through the pore of the channels is known as the “selectivity filter” and is responsible for discriminating between the different ionic species in the cell.
—
Selectivity filters
The selectivity filter of K+ channels is a highly conserved feature that is characterized by a signature amino acid sequence (TVGYG) (Heginbotham et al., 1994). As can be seen in Fig.1 the selectivity filter forms the most constricted part of the conduction pathway. It is so constricted that in order for the K+ ion to pass through most of the waters forming the hydration shell of the ion have to be removed. In the selectivity filer these water molecules are replaced by polar oxygen atoms of the protein. In Fig.2 it can been seen that the selectivity filter is comprised of four distinct binding sites (S1-S4, not to be confused with the S1-S4 transmembrane helices of voltage-sensor domains) (Zhou et al., 2001).
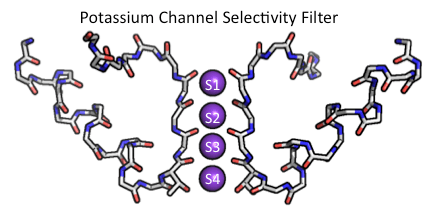
Figure 2. The KcsA potassium channel selectivity filter main chain structure (Zhou et al., 2001). The front and back subunits of the K+ channel tetramer as well as the side chains of the residues shown have been removed for clarity. The side chain of amino acid residue threonine of the signature sequence is shown since it participates in coordinating a K+ atom in site S4. Atoms are colored with carbon grey, oxygen red and nitrogen blue. This figure was made using PyMol.
It is via dehydration in these four biding sites that the selectivity for K+ is achieved (Zhou et al., 2001). In the absence of K+ the selectivity filter undergoes a conformational change making it non-conductive (Zhou et al., 2001; Zhou et al., 2003). Much work has been done to understand the exact mechanism of selectivity but my purpose here is just to introduce the idea and to show what an ion channel selectivity filter looks like. For more information and reviews on channel selectivity please see: Doyle et al., 1998; Gouaux & MacKinnon, 2005; Alam & Jiang, 2008; Ye et al., 2010; Roux et al., 2011.
The recent structures of the voltage-gated Na+ channels have given us our first look at the structure of a selectivity filter that selects for Na+ ions (Payandeh et al., 2011; Payandeh et al., 2012; Zhang et al., 2012). Surprisingly, the selectivity filter for the smaller Na+ ion is wider than that of the larger K+ ion (Fig.3). It is thought that the selectivity filter of Na+ channels may select for a hydrated Na+ ion as opposed to dehydrating the ion before conduction as in the case of the K+selectivity filter. However, the Na+ channel structures raises more questions than answers concerning Na+ selectivity and as always more experiments need to be performed.
No structure of a Ca2+ selectivity filter has been determined.
—
Conclusions
Lots of work has been done to understand how organic molecules are capable of discriminating between different ions in solution (Pedersen, 1988). Therefore, we know how small organic molecules, such as crown ethers and related ionophores, are able to discriminate between different ions. The major determining factors are size and charge (Fig.3) (Pedersen, 1988). Also the ratio of charge to size known as “charge density” plays an important role, for example, although Na+ and Ca2+ have very similar sizes, the charge density of Ca2+ will be twice as high since it has twice the charge. The charge density of an ion greatly impacts on the energetics of the interactions between the ion and other molecules such as water or proteins (Collins, 1997).
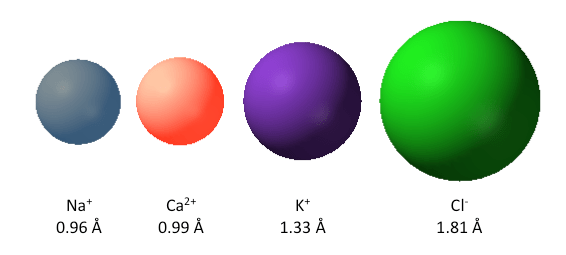
Figure 3. The relative sizes of the most prevalent inorganic ions of cells. The numbers represent the the ionic radii in Angstroms (10-10 m). Not shown is the proton which has a radius of 0.8775 fm (10-15 m).
Although these studies of small molecules have defined the priciples of ion selectivity we are still unsure how directly these same principles will apply to proteins. Clearly, size and charge matter, but further constraints on protein conformation and functionality may make ion selectivity less straight forward. For example, it is understood that highly selective ionophores can bind very tightly to their respective ion, however, in the case of channels tight binding would be counter productive. The role of channels is to conduct ions, therefore they have to be equally good at letting go of ions as they are at selecting them. This is a tough problem.
So where does Hv fit in?
Hv is an even more special case since it conducts protons (H+). Unlike the larger ions in Fig.3, H+ is not an atom, it is a particle. Compared to the ions in Fig.3, the size of the proton would not amount to a single pixel of the image, it is on the order of 100,000 times smaller than Na+ (check out the proton of wikipedia). Therefore the mechanism of H+ conduction and selectivity must be very different. There are a number of ideas floating around in the literature about how this might be accomplished, my next series of posts will discuss some of these ideas and explore what we know about how protons conduct through the Hv channel.
—
Works Cited and Further Reading
Alam, A., & Jiang, Y. (2008). Structural analysis of ion selectivity in the NaK channel. Nature structural & molecular biology, 16(1), 35–41. doi:10.1038/nsmb.1537
Collins, K. D. (1997). Charge density-dependent strength of hydration and biological structure Biophysj, 72(1), 65–76. doi:10.1016/S0006-3495(97)78647-8
Doyle, D. A., Morais Cabral, J., Pfuetzner, R. A., Kuo, A., Gulbis, J. M., Cohen, S. L., Chait, B. T., et al. (1998). The structure of the potassium channel: molecular basis of K+ conduction and selectivity Science, 280(5360), 69–77.
Gouaux, E., & Mackinnon, R. (2005). Principles of selective ion transport in channels and pumps Science, 310(5753), 1461–1465. doi:10.1126/science.1113666
Guo, M., Chong, Y. E., Shapiro, R., Beebe, K., Yang, X.-L., & Schimmel, P. (2009). Paradox of mistranslation of serine for alanine caused by AlaRS recognition dilemma Nature, 462(7274), 808–812. doi:10.1038/nature08612
Heginbotham, L., Lu, Z., Abramson, T., & Mackinnon, R. (1994). Mutations in the K+ channel signature sequence. Biophysical Journal, 66(4), 1061–1067. doi:10.1016/S0006-3495(94)80887-2
Long, S. B., Tao, X., Campbell, E. B., & Mackinnon, R. (2007). Atomic structure of a voltage-dependent K+ channel in a lipid membrane-like environment. Nature, 450(7168), 376–382. doi:10.1038/nature06265
Morais-Cabral, J. H., Zhou, Y, & Mackinnon, R. (2001). Energetic optimization of ion conduction rate by the K+ selectivity filter Nature, 414(6859), 37–42. doi:10.1038/35102000
Payandeh, J., Gamal El-Din, T. M., Scheuer, T., Zheng, N., & Catterall, W. A. (2012). Crystal structure of a voltage-gated sodium channel in two potentially inactivated states Nature, 486(7401), 135–139. doi:10.1038/nature11077
Payandeh, J., Scheuer, T., Zheng, N., & Catterall, W. A. (2011). The crystal structure of a voltage-gated sodium channel. Nature, 1–7. Nature Publishing Group. doi:10.1038/nature10238
Pedersen, C. J. (1988). The discovery of crown ethers Science, 241(4865), 536–540. doi:10.1126/science.241.4865.536
Roux, B., Bernèche, S., Egwolf, B., Lev, B., Noskov, S. Y., Rowley, C. N., & Yu, H. (2011). Ion selectivity in channels and transporters The Journal of General Physiology, 137(5), 415–426. doi:10.1085/jgp.201010577
Ujwal, R., Cascio, D., Colletier, J.-P., Faham, S., Zhang, J., Toro, L., Ping, P., et al. (2008). The crystal structure of mouse VDAC1 at 2.3 A resolution reveals mechanistic insights into metabolite gating Proceedings of the National Academy of Sciences of the United States of America, 105(46), 17742–17747. doi:10.1073/pnas.0809634105
Ye, S., Li, Y., & Jiang, Y. (2010). Novel insights into K+ selectivity from high-resolution structures of an open K+ channel pore. Nature structural & molecular biology, 17(8), 1019–1023. doi:10.1038/nsmb.1865
Zhang, X., Ren, W., DeCaen, P., Yan, C., Tao, X., Tang, L., Wang, J., et al. (2012). Crystal structure of an orthologue of the NaChBac voltage-gated sodium channel. Nature. doi:10.1038/nature11054
Zhou, Yufeng, & Mackinnon, R. (2003). The occupancy of ions in the K+ selectivity filter: charge balance and coupling of ion binding to a protein conformational change underlie high conduction rates. Journal of Molecular Biology, 333(5), 965–975.
Zhou, Y, Morais-Cabral, J. H., Kaufman, A., & Mackinnon, R. (2001). Chemistry of ion coordination and hydration revealed by a K+ channel-Fab complex at 2.0 A resolution Nature, 414(6859), 43–48. doi:10.1038/35102009
Comments
7 Responses to “Pores and Selectivity Filters”Trackbacks
Check out what others are saying...-
[…] “omega pathway” or “omega pore”. As I discussed in my previous post, Pores and Selectivity Filters, pores are aqueous pathways through membrane proteins that allow the diffusion of ions between the […]
Great introduction! I’ll definitely be following your blog for future posts.
Thanks Chris!
Interesting Stuff James.
Can you do a piece on their evolution? What protein was co-opted during the evolution of voltage gated channels? Did they evolve many times throughout the tree of life? Did they evolve once early on? I’m just curious… Understanding multiple origins could help to answer whether or not you should expect the mechanisms to be the same. It would also help to determine how easy it is for any pore to become voltage gated…
I don’t know… just wondering.
Hello Jav,
Thanks for the comment. I haven’t done too much research on the evolution of voltage sensing, but here are a few things that I know:
The different channels have evolved through a series of gene duplications
It is rare to find voltage-gated channels is prokaryotes (there are plenty of examples, most notably KvAP, but they are not nearly as ubiquitous as they are in eukaryotic cells) and the function of voltage-sensing in prokaryotes is poorly understood (they certainly don’t use them to pass signals between cells)
Hv is found in all eukaryotes that have been examined even the single celled ones where its role is central to maintaining pH homeostasis (I will definitely be writing more about this in the future)
If I had to guess I would imagine that Hv may have evolved first. It seems that as soon as cells evolve internal membranes (ie prkaryotic to eukaryotic transition) they got a strong selective advantage from being able to selectively conduct proton in and out of the different compartments. Metabolism is generally protonogenic (I think I just made that word up but you get the drift). This is pure speculation of course but the voltage sensor may have just evolved once and then been co-opted by pores to facilitate more complex signaling. In this way we would expect the mechanism of voltage-sensing to be related across the channels. However, evolution is really good at tweaking and adapting mechanism to suit different molecular niches. I agree that this would make an interesting future post, I will add it to the list, but clearly I need to do more reading.
Nice post, yet again.
Evolution of the Kv channel superfamily has captivated my interest for long. I don’t work on it and so cannot expand on it much but I’ll throw in my two cents as well.
A plethora of voltage-gated Kv channels are found in prokaryotes as well (all having the Kv selectivity filter sequence, the periodic Arg in S4, the conserved negative charges as well as the conserved phenylalanine). In addition a huge number of bacterial voltage-gated sodium channels (homotetrameric like NaChBac) are also annotated in the Swiss-Prot database (consensus selectivity sequence LESWAS). DE Clapham’s group has shown some evidence that in certain bacterium these Nav channel play an important role in chemotaxis/cell motility. Thus voltage-gated channels might actually have a role to play in bacteria survival, although cell-cell communication is unlikely the function of these proteins in bacteria. The existence of these proteins in bacteria must point towards an older, common ancestral history shared between the two proteins.
Earlier I thought that the pore domain and the VSD evolved independently and then fused together at one point in time and thereafter created a family of proteins which evolved as voltage-gated ion channels. The VSD and the pore are structurally independent (KvAP isolated VSD folds properly; the pore domain of several bacterial Nav channels have been cleaved off the full-length protein and shown to fold and function (done by Dan Minor’s group)). If this hypothesis about evolution is true, one should be able to find voltage-sensor only proteins like the CiVSP or the Hv channels in bacteria as well. However, the main problem is that although pore-domain only proteins in bacteria are plentiful, voltage-sensor only proteins currently have no known homologs in bacteria.
However, of late I made some observations which are beginning to make me think about this issue a bit differently. If you take a look at the crystal structure of LeuT (sodium coupled leucine transporter), solved by Eric Gouaux’s lab, and then squint your eyes and then open them again – you can unmistakably see a 4 helical bundle-like structure, very amusingly similar to the VSD. Additionally looking at the structure of the zinc transporter YiiP, you can see an arrangement of 6 helices which are strongly reminiscent of a single subunit of the Kv channel family. These curious structures and a cursory glance of ion channel evolution literature makes me think that the transporters were the mother proteins which at some point during evolution split into 2 fragments – one which evolved at the VSD and the other as the pore. Then a bit further down in the history of the earth, these fragments fused again and evolved as the VGICs.
Exploiting the evolutionary history of proteins to understand the principles of design and function is exciting and beautiful – as a community, I don’t think we have paid much attention to it. What is hot right now is finding bacterial homologs of important proteins and solving their structure – which is fantastic – but we can do so much more.
On a related known, very recently multiple voltage-gated sodium channels were cloned from sea anemone (invertebrates) and functionally expressed. These channels were peculiar as they did not have the standard DEKA locus selectivity filter but rather other filter sequences (eg. DEET, DEEA, DKEA, etc.) – the paper proposed a mechanism of evolutionary convergence of the sodium channel selectivity filter sequence and might have some useful information about the nature of evolution).
Thanks again. I really find your idea about the evolution of the VSD from an ancestral amino acid transporter very fascinating. They both have potential open-outward and open-inward conformations. Maybe it was an old arginine transporter.
Couldn’t agree more about how interesting and important the understanding of the evolution of these proteins is to our understanding of their mechanism. Also we still need more structures.